Multi measure plugin imagej9/9/2023  
However, the yellow and yellow-green actually have exactly the same color! You can verify this yourself by calling File › Open Samples › Spirals Macro in Fiji. Most people might think that the image contains 4 distinct colours: 2 sets of thin spirals are in dark red and dark green, and 2 thick prominent spirals of yellow-green and yellow. The problems with red/green merge images, aside from obvious issues with red-green colour-blind people, is that the perception of human eyes and brain can be fooled very easily. When considering colocalization, too often composite images of red and green channels are considered sufficient. If I want to show DAPI in green and EGFP as magenta, there is nothing “wrong” about that. 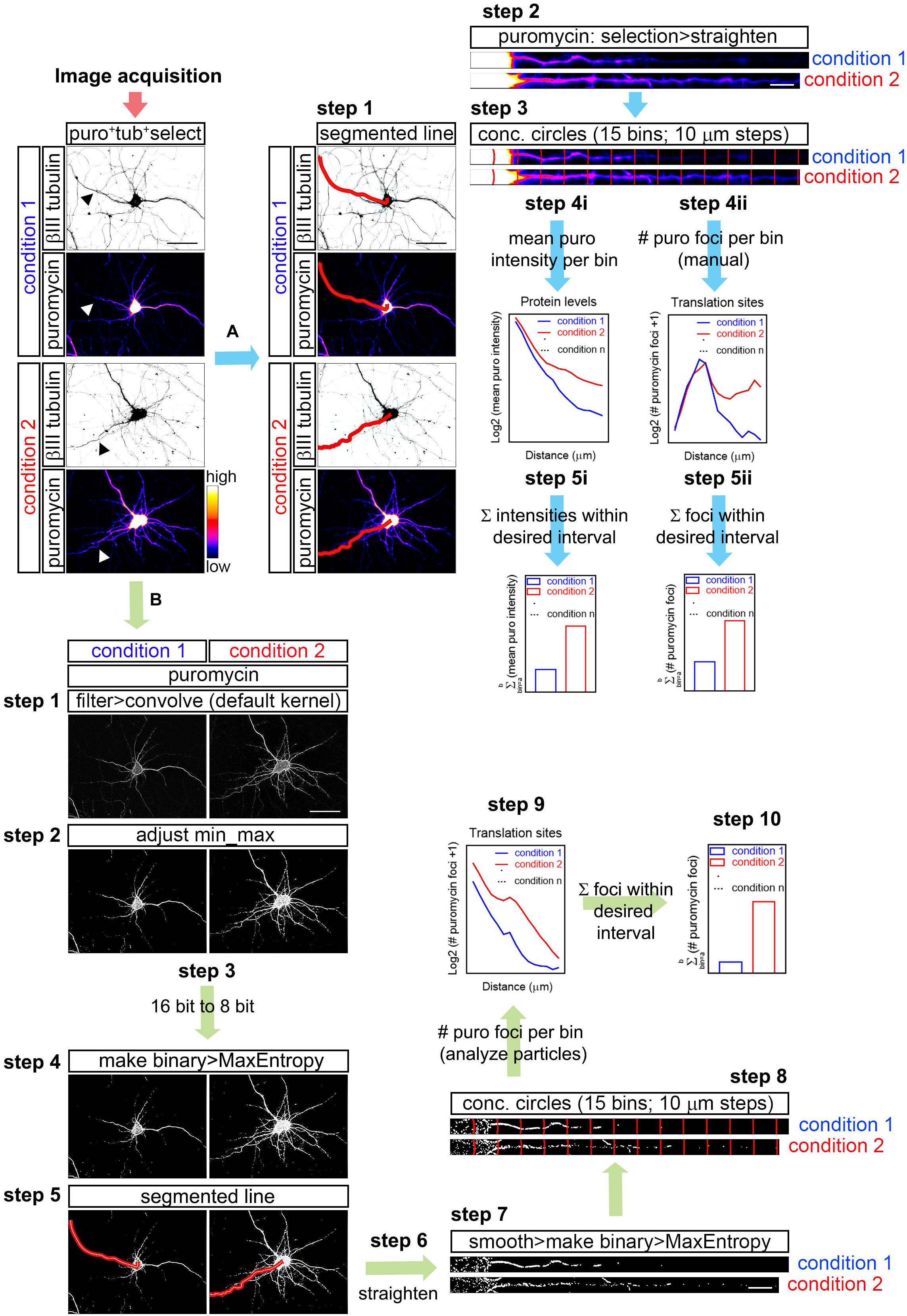 
These false colors are only useful to tell which channel is which. 
In the majority of fluorescence microscopy images, all the “colors” of a multi-channel image are captured using a monochromatic detector that doesn’t know what color the photons that hit it are that is determined by the fluorescence emission filters we use. You can find more details about optical resolution and image pixel spacing in the “Notes and precautions” section below. Additionally, the spatial resolution of your image must be sufficient to actually support your hypothesis. The colocalization measurement we make only means anything in relation to the spatial scale we are working at, so it needs to be explicitly stated. We must colocalise at some defined and explicit spatial scale: In our case the optical resolution or image pixel/voxel spacing, whichever is the larger value in nm, micrometers, mm, meters, km, etc. Practically, our situation lies between the two extremes. At the other extreme, a universe of one voxel (not cubic of course) is completely colocalised - everything is inside it. So, actually nothing is “really” colocalised. The Pauli exclusion principle states that two particles cannot have the same quantum numbers so they cannot be in the same place. For this reason, colocalization is most often used to determine if a protein is localizing to an organelle or other well defined cellular structure.įor more information on colocalisation and for how to correctly capture quantitative fluorescence microscopy images suitable for colocalisation analysis, look here: Image Processing Courses at BioDIP, Dresden. Regardless of your microscope, this volume is many, many times greater than the volume of a single protein. Importantly, colocalization results cannot indicate that two proteins/molecules are bound or interacting, only that they are both localized to within a certain volume, and is mostly dependent upon your microscope and its acquisition parameters. It could mean that one signal of one channel is contained within the bounds of another, or that your stains/dyes are typically found separated by a certain distance or are generally clustered, or simply that the signal from both channels overlap each other when imaged at a particular spatial resolution. The specific nature of that correlation, and what it means for your research, can vary quite a bit. Generally speaking, when we evaluate colocalization, we are usually attempting to demonstrate that a significant, non-random spatial correlation exists between two channels of a dual color image. First you have to define what you mean by colocalisation, and that is not trivial. Does anyone know of a solution within ImageJ, or alternatively another method such as through MATLAB?Įxample of two neurons, green circles indicate ROI position from day one scan, while current neuron position is offset to the right.Suppose you are given some images by a colleague, or have some images of your own, and you want to measure the amount of colocalisation between two of the dyes or stains in the images. This is necessary as I am generally dealing with hundreds of ROIs at once, making manually adjusting each ROI prohibitive. While Multi Measure does have a tool to update the position of individual ROIs, I cannot find (in ImageJ or on the internet) a way to shift all of my ROIs at once in the same direction. This means my ROI map from the first day does not line up with where the neurons are in the second day scan. Between days, my scans tend to be off from one another slightly in the XY coordinates. The plugin has been working wonderfully for this, however in more recent experiments I am repeatedly imaging from the same area over multiple days. I’ve been using the Multi Measure plugin to select ROIs and get the mean intensity of each ROI at each time point in the stack. My experiments involve two photon calcium imaging of neurons in vivo, which outputs data as a large time series of individual images (stack). 
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
